- Repository · LinkedIn · Tweet · BlueSky
- ⸻ 2026-03-04
A data lakehouse for biology’s sparse measurements  ¶
¶
One avenue into the future of biotech is scaled learning from multi-modal datasets. Given that the union of these datasets can easily span millions of sparse features, they can’t be queried through any established data infrastructure. Warehouses are too rigid, data lakes can’t be queried, and tabular lakehouses don’t understand the formats. Biology needs a data lakehouse with support for bio-formats and registries.
Multi-modal datasets¶
Early discovery biotech often feels like stumbling around a dark room with a flashlight: Each assay and experiment provides a narrow slice of insight into a highly complex system with more exceptions than rules. Multi-modal data such as transcriptomics, proteomics and high-content imaging gives researchers additional spotlights, expanding their field of view. But to understand what they’re seeing, they need to stitch these data modalities together into a single coherent whole. Most biological data consists of large numbers of relatively small datasets from different experiments, assays, labs and protocols. Some are large sparse matrices, such as single-cell gene expression data, with additional annotations on both rows and columns. Others, such as sequences or image features, are more conventional tables with a small number of columns. Most are annotated with complex, external ontologies or vocabularies linked to additional structured and unstructured metadata. And even datasets that are purportedly the same “type” often have slightly different schemas. Because of the partially overlapping features between these datasets, they conceptually fit together into a giant, sparse “feature matrix” where each “row” is an observation and each “column” a numerical measurement, a measurement of a discrete/categorical entity, textual records, or tensor data (for example, in the case of images or embeddings).

To build multi-modal machine learning models, computational biologists need to extract data from this (conceptual) matrix. But because datasets are typically stored as files scattered across different directories and storage locations, doing this in practice is complex and time-consuming under the best of circumstances. In many circumstances, it makes training ML models across the theoretically available datasets practically impossible.
Data lakes and warehouses¶
General-purpose data storage tools and infrastructure weren’t built for this reality of biological data. Until recent years, the gold standard for managing data at scale was the data warehouse, which extends the paradigm of the classical relational database. But while data warehouses have no problem managing the scale of multi-modal biological data, the problem is the relational paradigm. Data warehouses expect data to fit into a fixed number of tables with consistent schemas and with new data appended below existing rows, as on the left in the figure below. This works well for data that is collected incrementally through a consistent process. But for batches of biological data from a constantly changing collection of experimental protocols and objectives, it doesn’t work well.

The data lake architecture was introduced for contexts like this that don’t provide enough consistency to use a data warehouse. Data lakes give users complete flexibility to store data of any type in any format. This flexibility has made data lakes the standard solution for multi-modal biological data. However, the only way data lakes are able to provide this level of flexibility is to completely eliminate structure and consistency. Each dataset becomes its own isolated universe of information, as in the center of the above figure. If you want to connect it to an external ontology or even another dataset in the same format, it’s a completely manual process. This is part of why computational biologists spend half their time on manual cleaning and integration work.
The Lakehouse¶
Data warehouses impose too much structure for multi-modal biological data. Data lakes don’t provide enough. So an intermediate framework, the data lakehouse, was introduced to find a better balance for exactly this kind of situation. As shown on the right of the previous figure, a data lakehouse functions as a layer on top of a data lake that records structural information about each isolated dataset that can be used to dynamically extract consistently formatted information and integrate it with other datasets, external ontologies and other resources.
This option provides the best of both worlds: enough flexibility to store data from any assay/experiment/protocol that might come up, with enough structure to enable intuitive querying and model training while eliminating manual cleaning and bookkeeping.
One specific implementation that’s based on integrating biological registries with support for biological data formats is put forward in the open-source tool lamindb, and illustrated in the figure below.

You can pip install lamindb or check it out on GitHub: laminlabs/lamindb.
Materials¶
The “1 million features” pencil drawing that, over the years, led to the first figure in this post.
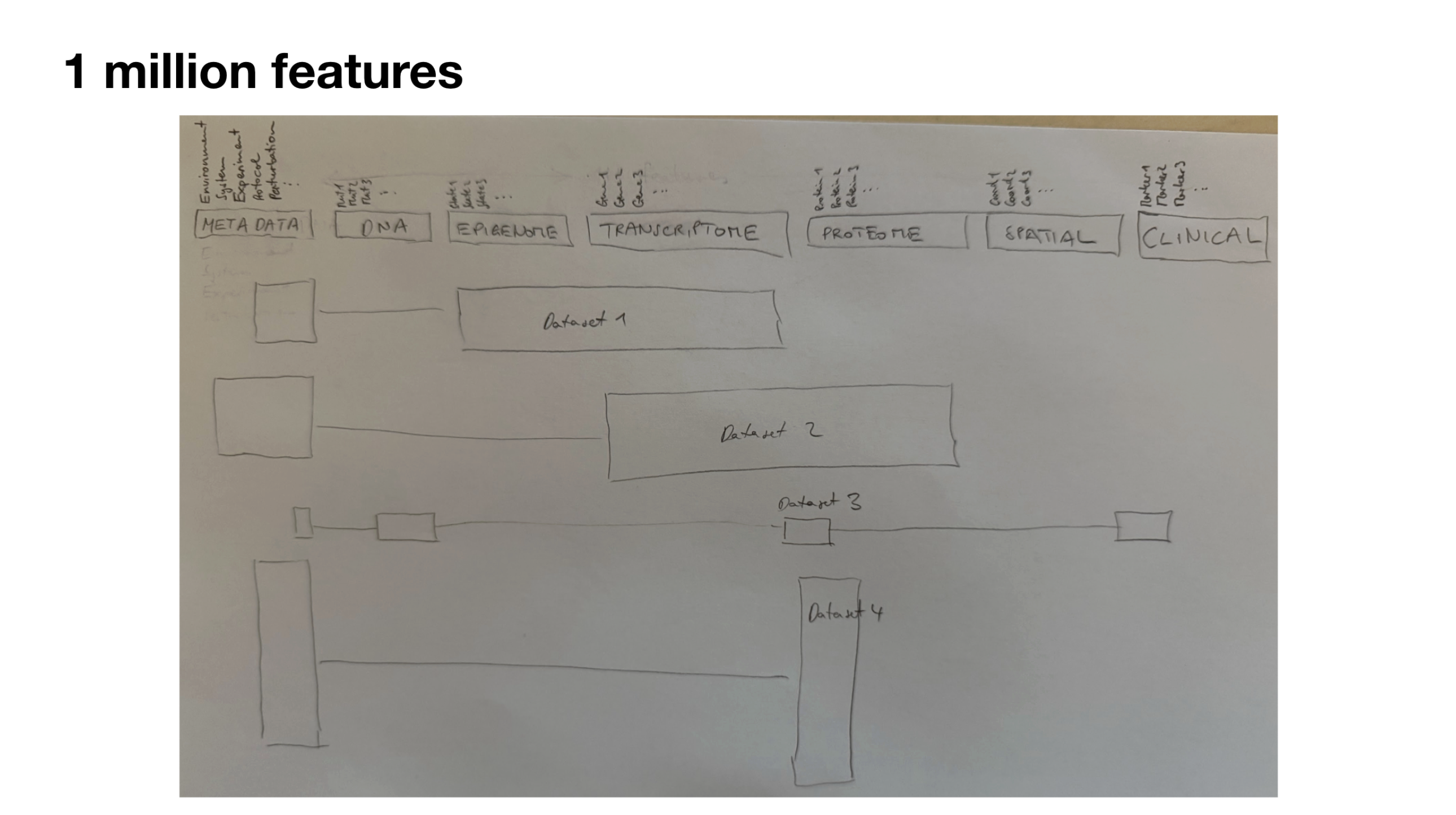
Background¶
Lamin engaged Jesse to explain the connection between biology’s “sparse measurements” and the lakehouse concept. This blog post is the result of that engagement.