- Database · Repository · LinkedIn · Tweet
- ⸻ 2026-05-12
Re-engineering the PerturBench benchmarking tasks with data lineage  ¶
¶
PerturBench (Wu, Wershof, Schmon, Nassar, Osinski, Eksi, Yan, et al., NeurIPS 2025) is a framework for benchmarking machine learning models that predict cellular transcriptional response to perturbations. Its core contributions are benchmarking tasks in the form of curated datasets and definitions of metrics, which are available from GitHub and Hugging Face, albeit without data lineage. To make it easy to see how exactly each dataset came about and assess model performance in light of that context, we re-ran all curation workflows using lineage tracking. We exemplify model training and evaluation, and show equivalence of the re-curated datasets with the originally deposited datasets.
While the situation has been improving in recent years through efforts like PerturBench[1], published scRNA-seq-based models have often been evaluated on inconsistent benchmarks, making it hard to know what works and going counter to the fact that machine learning breakthroughs need well-curated datasets and well-defined tasks. PerturBench is one of several efforts in the field and was preceded by an Open Problems benchmark, published at NeurIPS 2024[2]. Another example for a similar benchmarking effort is last year’s Arc Virtual Cell Challenge. For a video introduction to PerturBench, you can watch this episode of Valence Labs’ MultiOmics Reading Group.
The NeurIPS PerturBench submission hosts its six curated benchmarking datasets on Hugging Face. These files, however, don’t reveal how the curation was done.
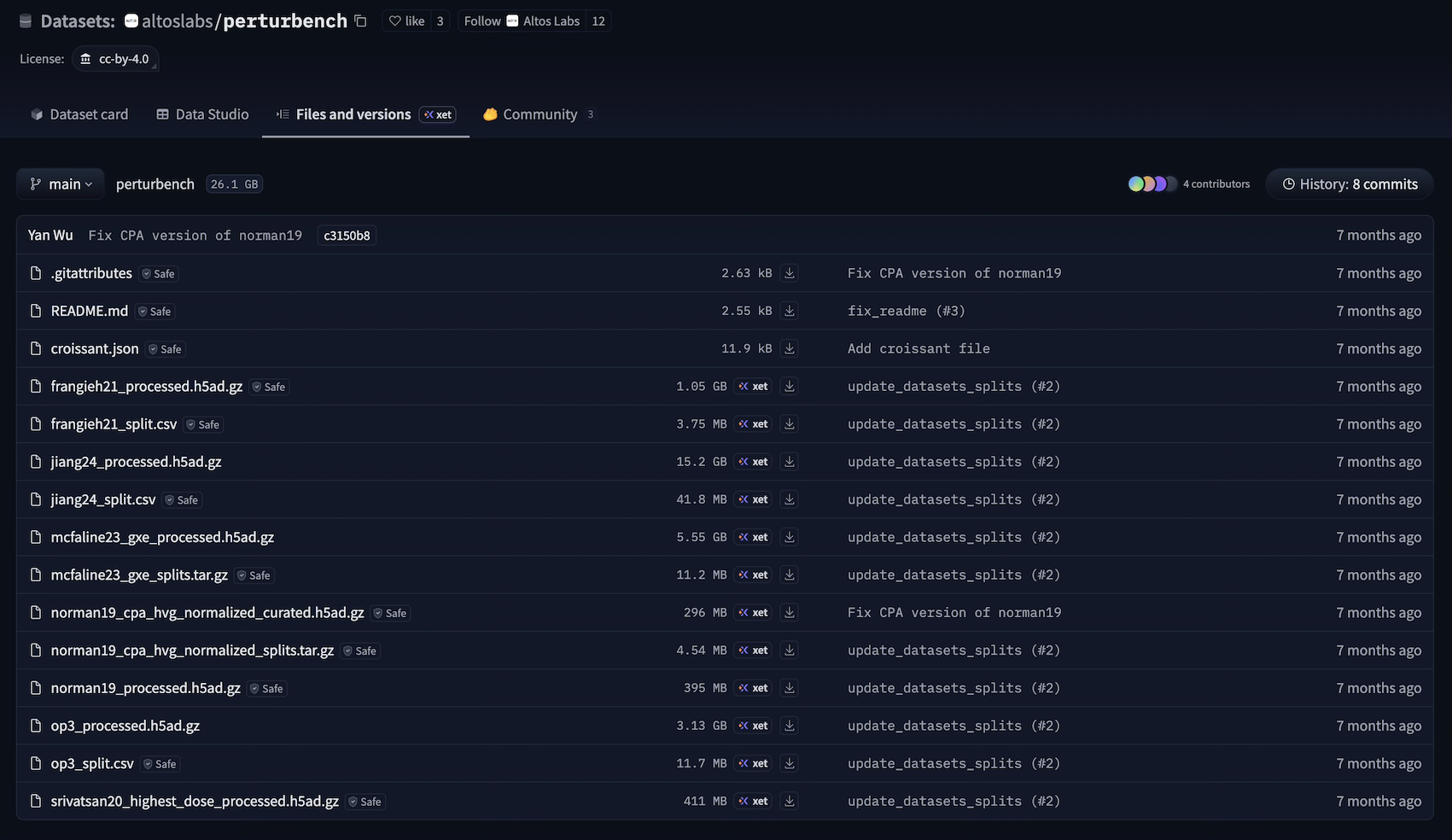
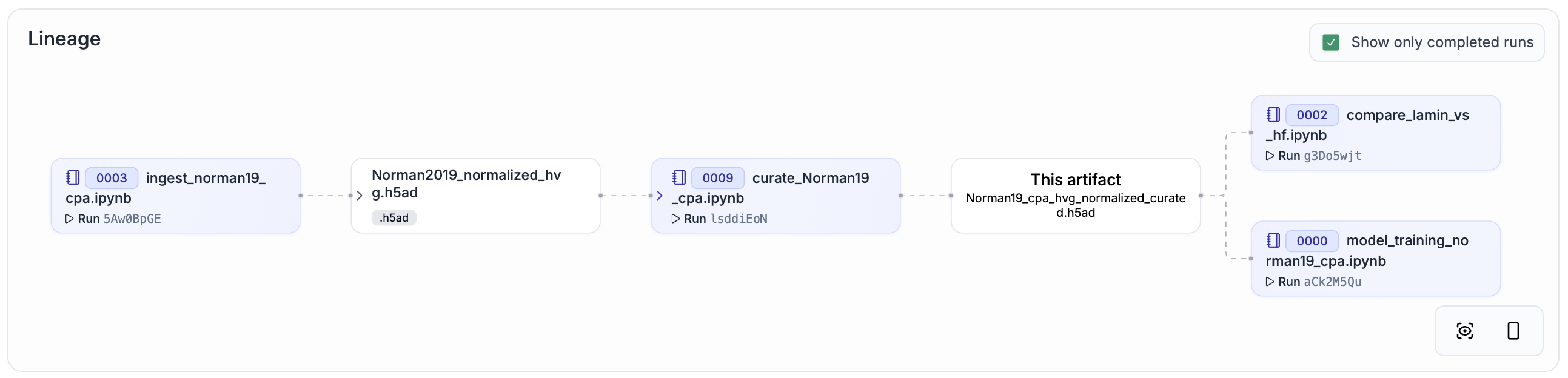
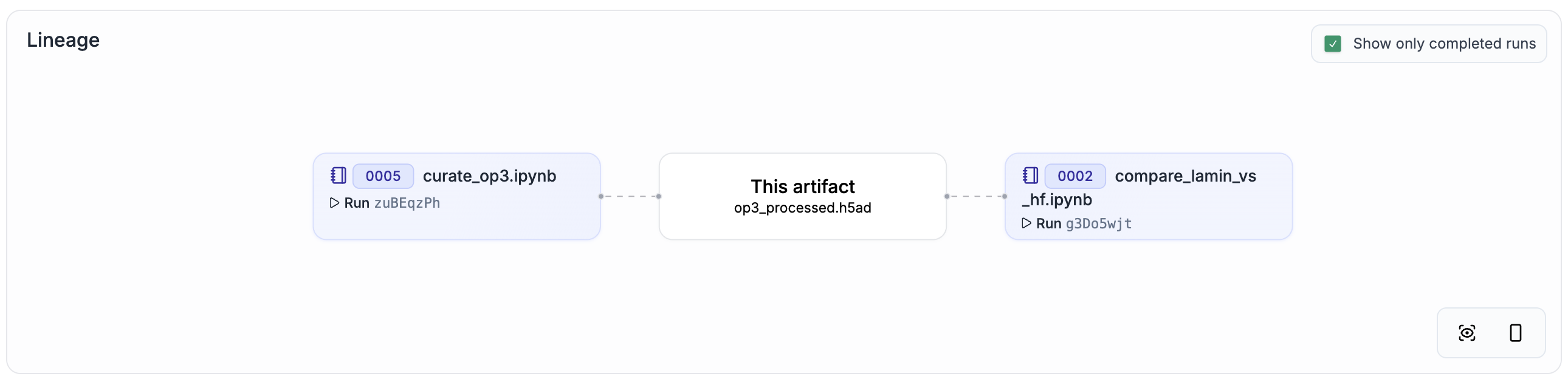
To make data lineage easy to browse and understand, we re-ran all curation steps with ln.track() added to the source code. You can explore the result by clicking on the link in the “Dataset + lineage” column of the following table.
Reference |
Perturbation |
N |
Dataset + lineage (click the image to explore) |
|---|---|---|---|
Norman19[3] |
Genetic |
91,168 cells |
|
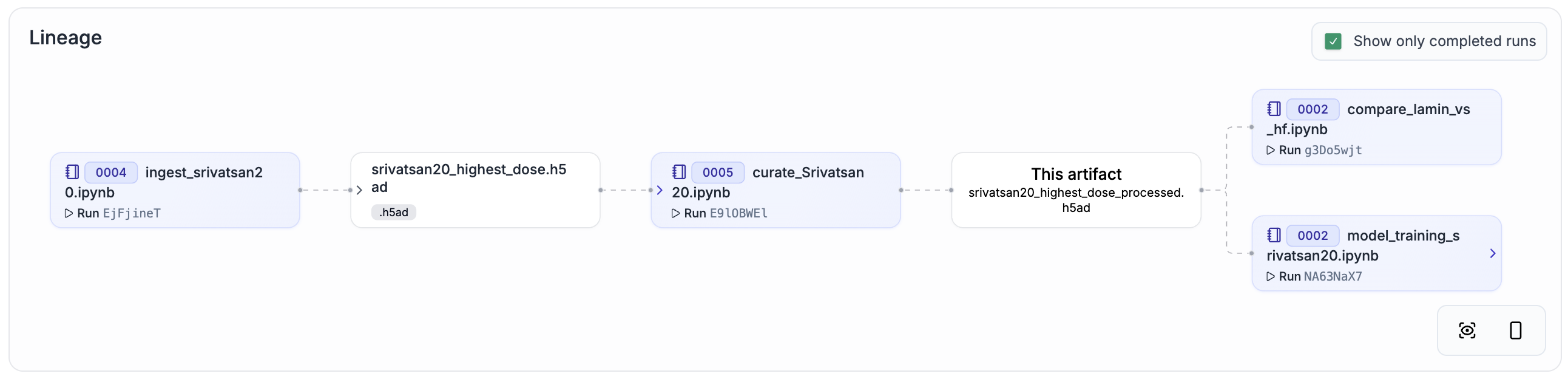
Srivatsan20[4] |
Chemical |
178,213 cells |
|
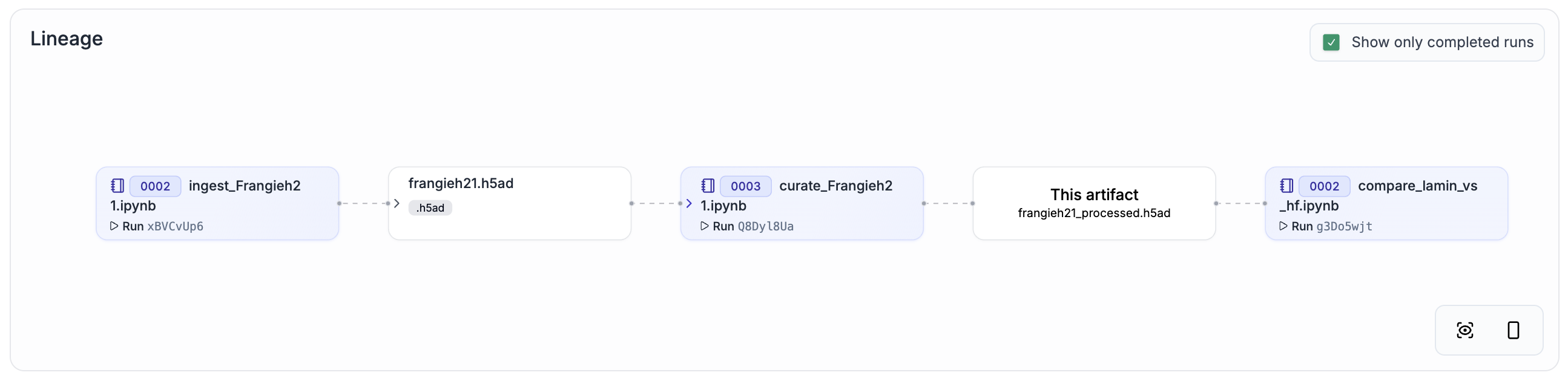
Frangieh21[5] |
Genetic |
218,331 cells |
|
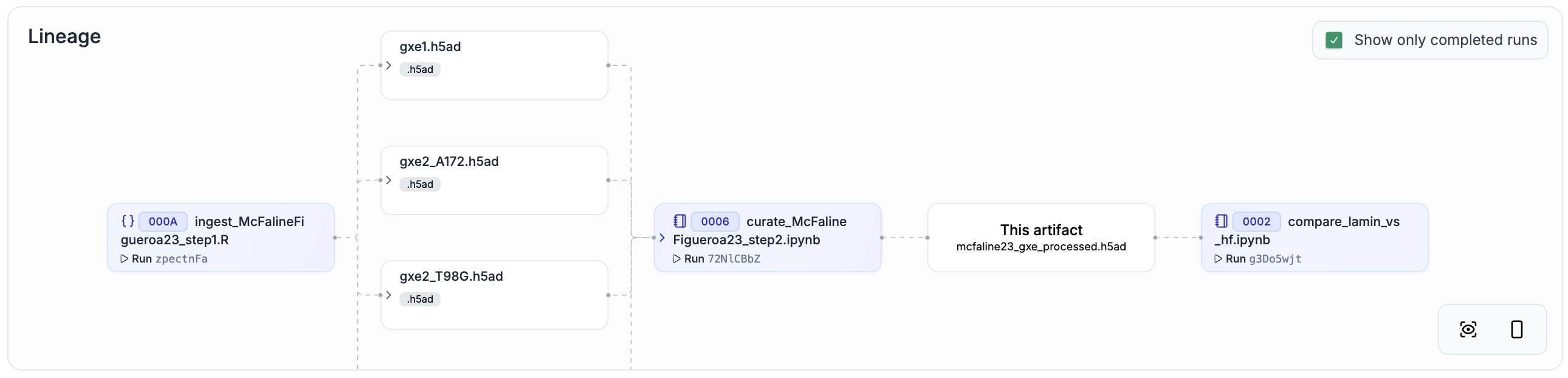
McFaline24[6] |
Genetic |
892,800 cells |
|
Jiang25[7] |
Genetic |
1,628,476 cells |
|
Szalata24 (OP3)[2] |
Chemical |
298,087 cells |
On a high level, the data flow is:
Raw data ingestion: Each dataset has a dedicated notebook (prefixed with
ingestion_) that ingests raw datasets.Curation: Curation notebooks (prefixed with
curate_) handle format conversion, preprocessing, metadata harmonization, and split generation.Training and eval: Curated datasets are loaded to train and evaluate models using the
PerturBenchPython framework.
For a comparison that shows the equivalence of the original datasets and the re-curated datasets, explore altoslabs/perturbench/transform/3bZAUr0kXokI. For a full training and model evaluation run, explore altoslabs/perturbench/transform/9dPgCiisCm1w.
This post was motivated by the desire to reproduce PerturBench’s training and eval results in a file-centric manner, omitting the detailed modeling of perturbational & biological metadata. Modeling and validating perturbations will be the topic of an upcoming post.
Code & data availability¶
Database: lamin.ai/altoslabs/perturbench. Repo: github.com/altoslabs/perturbench.
How to cite¶
Jain I, Namsaraeva A, Sun S, Wu Y & Wolf A (2026). Re-engineering the PerturBench benchmarking tasks with data lineage. Lamin Blog. https://blog.lamin.ai/perturbench